Project Due: May 11, 2020 at 11:59pm
Late Policy: Up to two slip days can be used for the final submission.
Please submit all required documents to CMS.
This is a partner project. You can either work alone, or work ONLY with your chosen partner. Failure to adhere to this rule (by e.g. copying code) may result in an Academic Integrity Violation.
Overview: In this project, you will be evaluating the impacts of memory allocation and parallelism on a learning algorithm. Since we are close to the end of the semester, I have planned this project to be relatively short so that you can finish it within one week, but please don't use this as a reason to put off starting on the project until the last day!
Instructions: This project is split into three parts: the implementation and evaluation of a version of SGD that doesn't allocate memory in its inner loop, the evaluation of the performance of the built-in multithreading capabilities of numpy, and the implementation of a multi-threaded version of SGD in Python.
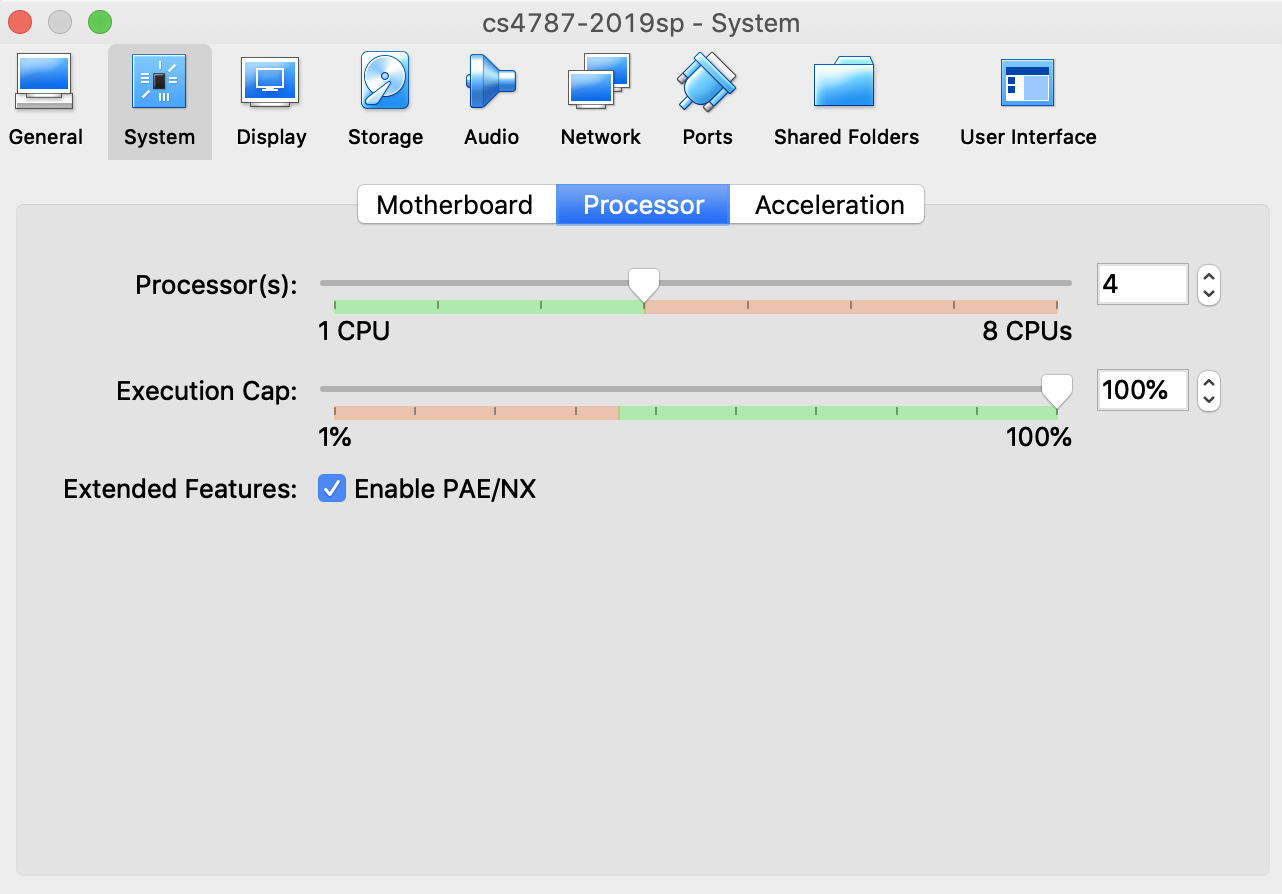
Note that for the multi-threading section of the assignment, you will need to run on a CPU with more than one core to get interesting results. If you are running on a VM, you will need to make sure that it is set up to allow for more than one thread of execution. To do this, you may need to increase the number of cores available in the VM settings, as illustrated in the figure below.

Note: There is no autograder for this project, since you can verify that your code outputs the correct thing by just comparing it to the baseline methods provided. Grading will be performed based on your lab report and on manual inspection of your code.
Also note: The experiments in this project can take some time to complete. On my computer, they take about 7 minutes to run in total (across all experiments). Please be sure to leave enough time to complete the experiments.
Part 1: Memory.
Background: Most numpy operations by default allocate a new array in which to return their result.
This extra allocation can be costly, and can cause harmful memory effects that slow down computation.
Fortunately, numpy has a mechanism that lets us avoid this.
Most numpy operations have an optional argument out which lets you specify an already-allocated array into which the result of the operation will be written.
This lets us perform a numpy operation without allocating any new memory.
For an algorithm like SGD, this means that we can perform all the allocations we need before entering the inner loop, and then just do all the computations in the inner loop with allocation-free numpy operations.
This can speed up the execution of the loop, as we'll see here. (For this part of this assignment, you should run experiments with the number of threads set to 1.)
sgd_mss_with_momentum (SGD + momentum with minibatching and sequential scan) from Programming Assignment 3 to return only the model arrived at at the end of all the iterations (rather than returning a list of models produced every so often).sgd_mss_with_momentum_noalloc that runs the same computations as your sgd_mss_with_momentum function, but avoids any allocations in the inner loop by pre-allocating any needed arrays.
out parameter that is a pre-defined array.Xs[:,ii] where Xs is a numpy array). If you want to use a slice, pre-allocate it outside the inner loop by running something like numpy.ascontiguousarray(Xs[:,ii]), which allocates that slice as a dense array.sgd_mss_with_momentum function.sgd_mss_with_momentum and sgd_mss_with_momentum_noalloc) on the MNIST dataset using the following hyperparameters:
Part 2: Parallelism in Numpy.
os.environ["OMP_NUM_THREADS"] = "n"
os.environ["MKL_NUM_THREADS"] = "n"
os.environ["OPENBLAS_NUM_THREADS"] = "n"
n is replaced by the number of threads you want to use (some number larger than 1).
You will need to restart python to make this change.
Part 3: Multithreading in Python. So far we have looked at parallelizing implicitly using numpy multithreading (supported on top of OpenMP and controlled by those environment variables). There is another way we can take advantage of the multi-core capabilities of our CPU: to explicitly use python multithreading capabilities. This is what we will do in this part.
sgd_mss_with_momentum_threaded that runs the same computations as your sgd_mss_with_momentum_noalloc function, and uses python multithreading functionality from the threading standard library. Your function should use multithreading to parallelize across subsets of the minibatch. That is, if you are running on \(M\) machines with a minibatch of size \(B\) where \(B = M \cdot B'\), then to compute the minibatch SGD update
\[
W_{t+1} = W_t - \alpha \cdot \frac{1}{B} \sum_{m=1}^M \sum_{b = 1}^{B'} \nabla f_{i_{m,b,t}}(W_t).
\]
I have provided some starter code that gives a skeleton for an implementation that does the following.
sgd_mss_with_momentum_threaded on the MNIST dataset using the following hyperparameters:
Part 4: The Effect of Precision.
In class, we talked about how using lower-precision arithmetic can improve the throughput of machine learning applications. Here, we will test whether this is true for this training task. Numpy uses 64-bit floating point arithmetic by default: we will see here whether using 32-bit floating-point arithmetic (which is the standard precision used for most ML applications and ML frameworks) can improve the training speed.
sgd_mss_with_momentum_noalloc_float32 and sgd_mss_with_momentum_threaded_float32, which work the same as the similarly named functions you implemented earlier, except that they do their computations using 32-bit floating point numbers. You should do this by copying your implementations from Parts 1 and 3, and adapting them to use single-precision floats. You may find the numpy datatype numpy.float32 and the dtype argument to be useful here.dtype=numpy.float32 and sometimes .astype(numpy.float32) as needed.sgd_mss_with_momentum_noalloc_float32 using implicit numpy multithreading (as in Part 2) on the MNIST dataset using the following hyperparameters:
What to submit:
main.py.Setup:
pip3 install -r requirements.txt to install the required python packages