Professor Ron Elber
Cornell University
Department of Computer Science
4130 Upson Hall, Ithaca, NY 14853
Email: ron@cs.cornell.edu
Phone: (607) 255-7416
FAX: (607) 255-4428
General Field of Investigation:
Structure and dynamics of biological
molecules: Theory, algorithm developments and computer simulations. Software for
molecular dynamics simulation -- MOIL
and for protein recognition -- LOOPP
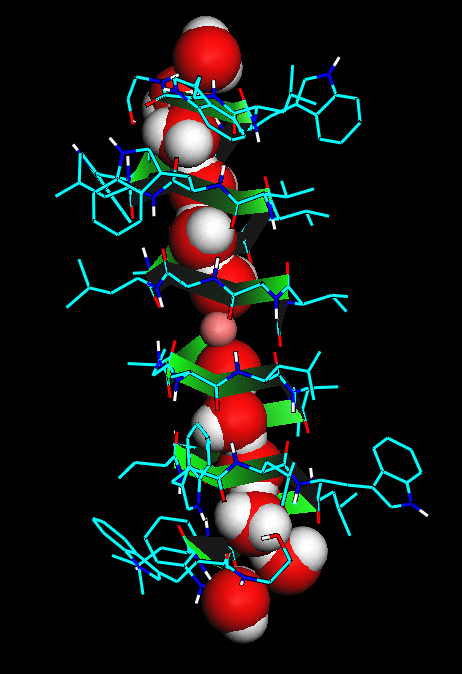
Figure: The gramicidin ion channel (sticks
and green ribbon) and a "permion" transported through it. A permion is a
collective "quasi-particle" that consists of about 8 water molecules (red balls
with white spots attached) and a sodium ion (the small pink ball in the middle) that moves
coherently (like in a queue) through the channel. A reaction path algorithm was
implemented into the Molecular Dynamics package MOIL [9] and was used to study the
properties of the permion. The picture was made with the program MOIL-View written by
Carlos Simmerling [10].
Topics of Research:
Ron Elber’s research focuses
on algorithm development and computer simulations of the structure and dynamics of
biomolecules.
Currently active research directions include
- The design and application of protein folding potentials.
Linear programming techniques are used to find optimal sets of parameters that recognize
the native folds of proteins. Potentials are designed for maximum accuracy and optimal
computational efficiency. We are studying a hierarchy of potentials, some functional forms
are trained to the highest accuracy and other formulations are less accurate but are
trained to search more rapidly for plausible conformations.
- The design and application of algorithms to compute long
time dynamics of biological molecules. Algorithms that are based on optimization of
functionals are investigated as an alternative to the usual approach of solving initial
value differential equations. Approximate stable trajectories of very broad time scales
can be obtained. Time steps larger by a factor of million compared to the common
simulation algorithms were used successfully.
- Mean-field global-optimization as a tool to determine
peptide structures in solution. We developed and applied mean field simulation techniques
that make it possible to compute peptide conformations in explicit solvent environment.
Structures of peptides of lengths smaller than ten amino acids were determined and
compared favorably with experiment.
- Searching for plausible diffusion pathways from buried
active sites. In a number of proteins (e.g. the globins) the active site is buried in the
protein interior and is not accessible to a solvated ligand. We developed efficient search
strategies for plausible diffusion pathways and gateways, which are induced by thermal
fluctuations of the protein. The search method, the Locally Enhanced Sampling approach
(LES), enables rapid exploration of small fraction of space which is of prime interest
(e.g., the accessible volume at the active site).
- Reaction path studies of slow processes in proteins. We
introduced novel computational approaches that make it possible to calculate minimum
energy paths for complex processes with motions of thousands of atoms, like the R to T
transition in hemoglobin, or ion permeation through a membrane channel. Statistical
approaches, which are based on the availability of the reaction paths, are finally used to
estimate the time scales measured experimentally.
Recent Representative Publications:
- C. Keasar, D. Tobi, R. Elber and J.
Skolnick, "Coupling the folding of homologous proteins", Proc. Natl. Acad. Sci.
USA, 95,5880-5883(1998)
- O. Schueler-Furman, Ron Elber and Hanah
Margalit, "Knowledge-based structure prediction of MHC class I bound peptides: A
study of twenty-three complexes", Folding and Design, 3,549-564(1998).
- J. Meller and R. Elber, Computer
simulations of carbon monoxide photo-dissociation in myoglobin: structural interpretation
of the B states. Biophys. J. 74,789-802 (1998)
- D. Mohanty, R. Elber and Devarajan
Thirumalai, Dimitrii Beglov and Benoit Roux, Kinetics of peptide folding: Computer
simulations of SYPFDV and peptide variations in water., J. Mol. Biol., 272, 423-442
(1997).
- C. Keasar, R. Elber and J. Skolnick,
Simultaneous and coupled energy optimization of homologous proteins: A new tool for
structure prediction, folding and design, 2,247-259 (1997)
- R. Olender and R. Elber, Calculation of
classical trajectories with a very large time step: formalism and numerical examples, J.
Chem. Phys., 105,9299-9315 (1996)
- A. Roitberg, R.B. Gerber, R. Elber and M.A.
Ratner, Anharmonic wave functions of proteins: quantum self-consistent field calculations
of BPTI. Science, 268,1319-1322 (1995).
- R. Elber, D. Chen, D. Rojewska and R.S.
Eisenberg, "Sodium in gramicidin: An example of a permion", Biophys. J.,
68,906-924(1995)
- R. Elber, A. Roitberg, C. Simmerling, R. Goldstein, H. Li,
G. Verkhivker, C. Keasar, J. Zhang and A. Ulitsky "MOIL: A program for simulations of
macromolecules", Computer Physics Communications, 91,159-189(1995)
- C. Simmerling, R. Elber and J. Zhang, "Moil-View - a
program for visualization of structure and dynamics of biomolecules and STO - a program
for computing stochastic paths", in "The proceeding of the Jerusalem symposium
on theoretical biochemistry", Modeling of Biomolecular Structures and Mechanisms,
1995, Ed. A. Pullman et al, 241-265, 1995 (Kluwer Academic Publishers, Netherlands)
Group Members:
- Dr. Alfredo Cardenas
- Dr. Avijit Ghosh
- Dr. Tamara Galor Naeh
- Dr. Octavian Teodorescu
- Dr. Harry Stern
Any
Questions or bug reports regarding this page should go to croft@cs.cornell.edu