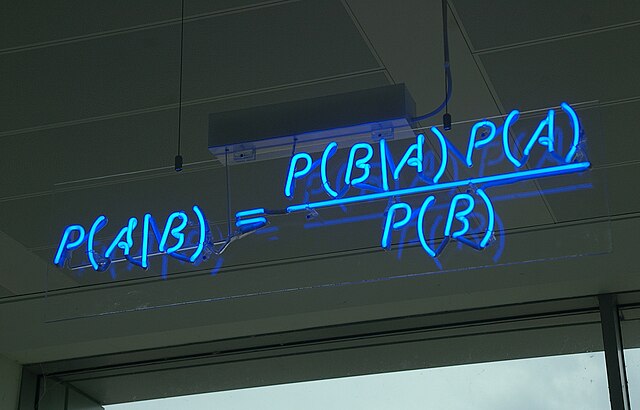
Recall the Bayes Optimal classifier. If we are provided with the true source distribution \( \mathcal{P} \) over example-label pairs \( (X, Y) \), we can predict the most likely label for \(\mathbf{x}\): \(\operatorname*{argmax}_y \mathcal{P}(y|\mathbf{x}). \) One very general approach to learning motivated by this is to estimate \( \mathcal{P} \) directly from the training data. If it is possible (to a good approximation) to construct an estimate \( \hat{\mathcal{P}} \approx \mathcal{P} \) we could then use the Bayes Optimal classifier on \( \hat{\mathcal{P}} \) in practice.
Many supervised learning strategies can be viewed as estimating \( \mathcal{P} \). Generally, they fall into two categories:
Suppose you find a coin and it's ancient and very valuable. Naturally, you ask yourself, "What is the probability that this coin comes up heads when I toss it?" You toss it \(n = 10\) times and obtain the following (ordered) sequence of outcomes: \(D=\{H, T, T, H, H, H, T, T, T, T\}\). Based on these samples, how would you estimate \(P(H)\)? We observed \(n_H=4\) heads and \(n_T=6\) tails. So, intuitively, $$ P(H) \approx \frac{n_H}{n_H + n_T} = \frac{4}{10}= 0.4. $$ But is this reasoning valid? Can we derive this more formally?
The estimator we just mentioned is the Maximum Likelihood Estimate (MLE). For MLE you typically proceed in two steps.
Let us return to the coin example. A natural assumption about a sequence of coin tosses (for a particular coin with a fixed probability of coming up heads) is that the results of the flips are independent. Formally, if \( X_i \in \{0,1\} \) is the indicator random variable representing the event that the result of the \(i\)th flip is heads, and we suppose that the probability of the coin coming up heads is \( \theta \in [0,1] \), then \[ \mathbf{P}(X_i = x_i; \theta) = \begin{cases} \theta & \text{ if } x_i = 1 \\ 1 - \theta & \text{ if } x_i = 0 \end{cases} = \theta^{x_i} (1 - \theta)^{1 - x_i}. \] By independence, the probability of many such flips coming up in any particular way (i.e. in some specific order \( x_1, x_2, \ldots, x_n \)) will be the product of the probabilities \begin{align*} \mathbf{P}(X_1 = x_1, X_2 = x_2, \ldots, X_n = x_n; \theta) &= \prod_{i=1}^n \mathbf{P}(X_i = x_i; \theta) \\&= \prod_{i=1}^n \theta^{x_i} (1 - \theta)^{1 - x_i} \\&= \theta^{\sum_{i=1}^n x_i} (1 - \theta)^{\sum_{i=1}^n (1 - x_i)}. \end{align*} We can write this a little more consisely if we let \( n_H = \sum_{i=1}^n x_i \) denote the number of heads and \( n_T = \sum_{i=1}^n 1 - x_i \) denote the number of tails. In this case, we would just have \[ \mathbf{P}(X_1 = x_1, X_2 = x_2, \ldots, X_n = x_n; \theta) = \theta^{n_H} (1 - \theta)^{n_T}. \]
MLE Principle: Find \(\hat{\theta}\) to maximize the likelihood of the data, \( \mathbf{P}(\mathcal{D}; \theta)\): \begin{align} \hat{\theta}_{MLE} &= \operatorname*{argmax}_{\theta} \, \mathbf{P}( \mathcal{D} ; \theta), \end{align} where here we slightly abuse notation to let \( \mathcal{D} \) also denote the event that the dataset \( \mathcal{D} \) is observed.
Often we can solve this maximization problem with a simple three step procedure:
Returning to our coin-flipping example, we can now plug in the definition and compute the log-likelihood: \begin{align} \hat{\theta}_{\operatorname{MLE}} &= \operatorname*{argmax}_{\theta} \; \mathbf{P}(D; \theta) \\ &= \operatorname*{argmax}_{\theta} \; \theta^{n_H} (1 - \theta)^{n_T} \\ &= \operatorname*{argmax}_{\theta} \; \log\left( \theta^{n_H} (1 - \theta)^{n_T} \right) \\ &= \operatorname*{argmax}_{\theta} \; n_H \cdot \log(\theta) + n_T \cdot \log(1 - \theta). \end{align} We can then solve for \(\theta\) by taking the derivative and equating it with zero. This results in \begin{align} 0 = \frac{n_H}{\theta} - \frac{n_T}{1 - \theta} \Longrightarrow n_H - n_H\theta = n_T\theta \Longrightarrow \theta = \frac{n_H}{n_H + n_T}. \end{align} A nice sanity check is that \(\theta\in[0,1]\).
Suppose you have a hunch that \(\theta\) is close to \( \frac{1}{2} \). But your sample size is small, so you don't trust your estimate. Simple fix: Add \(m\) imaginary throws that would result in the \(\theta\) we have a hunch it is close to (e.g. \(\theta = 0.5\)). For example, we could add \(m\) Heads and \(m\) Tails to your data, which would result in the modified MLE estimate $$ \hat{\theta} = \frac{n_H + m}{n_H + n_T + 2m} $$ For large \(n\), this is an insignificant change. For small \(n\), it incorporates your "prior belief" about what \(\theta\) should be. But this was all very ad hoc. Can we justify this and derive it formally?
Model \(\theta\) as a random variable, with its own marginal distribution \( \mathbf{P}(\theta)\). Here \( \theta \) and the other observed variables \( (x_i, y_i) \) are jointly distributed.
Note that \(\theta\) is not a random variable associated with an event in a sample space. In frequentist statistics, this is forbidden. In Bayesian statistics, this is allowed and you can specify a prior belief \(\mathbf{P}(\theta)\) defining what values you believe \(\theta\) is likely to take on.
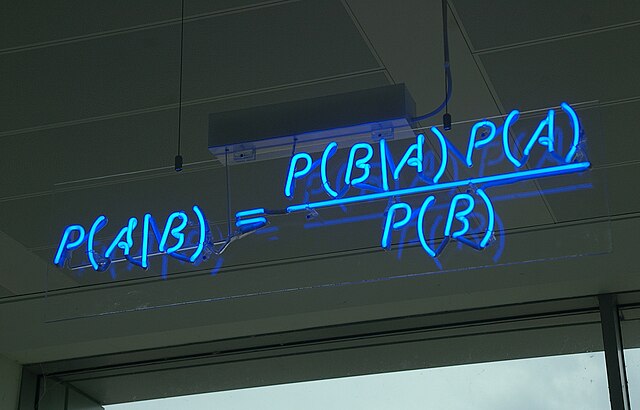
Now, we can look at \( \mathbf{P}(\theta \mid \mathcal{D} ) = \frac{ \mathbf{P}(\mathcal{D} \mid \theta) \mathbf{P}(\theta)}{ \mathbf{P}(\mathcal{D} )}\) (recall Bayes Rule!), where
A natural choice for the prior \(\mathbf{P}(\theta\)) is the Beta distribution: \begin{align} \mathbf{P}(\theta) = \frac{\theta^{\alpha - 1}(1 - \theta)^{\beta - 1}}{B(\alpha, \beta)} \end{align} where the beta function \[ B(\alpha, \beta) = \frac{\Gamma(\alpha) \Gamma(\beta)}{\Gamma(\alpha+\beta)} = \int_{0}^{1} \theta^{\alpha - 1} (1 - \theta)^{\beta - 1} \; d \theta \] is the normalization constant (if this looks scary don't worry about it, it is just there to make sure everything sums to \(1\) and to scare children at Halloween). Note that here we only need a distribution over a single random variable \(\theta\). (The multivariate generalization of the Beta distribution is the Dirichlet distribution.)
Why is the Beta distribution a good fit?
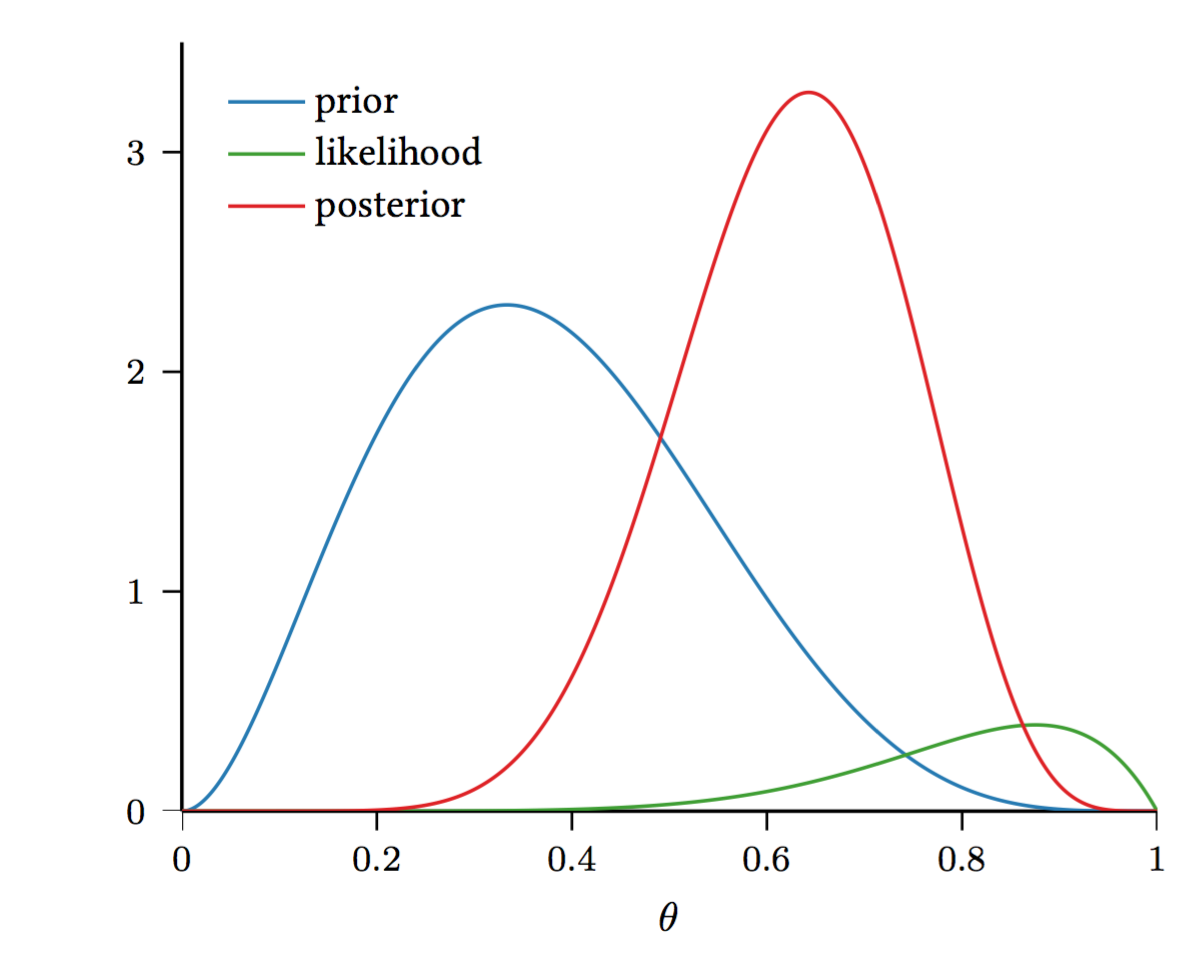
So far, we have a distribution over \(\theta\). How can we get an estimate for \(\theta\)?
A few comments:
MAP is only one way to get an estimator. There is much more information in \( \mathbf{P}(\theta \mid \mathcal{D})\), and it seems like a shame to simply compute the mode and throw away all other information. A true Bayesian approach is to use the posterior predictive distribution directly to make predictions about the label \(Y\) of a test sample with features \(X\). Here we consider \(X\) and \(Y\) to be a "fresh" test example independent of all the examples in the training dataset conditioned on \( \theta \). Concretely, this corresponds to the distribution over \( y \) given by \begin{align*} \hat{\mathcal{P}}(x, y) = \int \mathbf{P}(Y = y \mid X = x, \theta = \vartheta) \mathbf{P}(\theta = \vartheta | \mathcal{D} ) \; d\vartheta. \end{align*} Unfortunately, the above is generally intractable in closed form. Sampling techniques, such as Monte Carlo approximations, are used to approximate the distribution. A common exception where this is tractable is the case of Gaussian Processes.
Another exception is actually our coin toss example. Here, we don't really have features, so we can ignore the \( X \) in the above formulation. To make predictions using \(\theta\) in our coin tossing example, we can see that the distribution of a fresh flip is \begin{align} \hat{\mathcal{P}}(y) =& \int_0^1 \mathbf{P}(Y = y \mid \theta = \vartheta) \mathbf{P}(\theta = \vartheta | \mathcal{D} ) \; d\vartheta \\ =& \int_0^1 \mathbf{P}(Y = y \mid \theta = \vartheta) \cdot \frac{ \mathbf{P}(\mathcal{D} \mid \theta = \vartheta ) \mathbf{P}(\theta = \vartheta )}{\mathbf{P}( \mathcal{D} )} \; d\vartheta \\ =& \int_0^1 \vartheta^{y} (1 - \vartheta)^{1 - y} \cdot \frac{ \vartheta^{n_H + \alpha - 1} (1 - \vartheta)^{n_T + \beta - 1} }{B(\alpha, \beta) \cdot \mathbf{P}( \mathcal{D} )} \; d\vartheta. \end{align} If we let \( Z = B(\alpha, \beta) \mathbf{P}(\mathcal{D}) \), which is just a constant independent of \( y \), then \begin{align} \hat{\mathcal{P}}(1) =& \frac{1}{Z} \int_0^1 \vartheta^{n_H + \alpha} (1 - \vartheta)^{n_T + \beta - 1} \; d\vartheta \end{align} and \begin{align} \hat{\mathcal{P}}(0) =& \frac{1}{Z} \int_0^1 \vartheta^{n_H + \alpha - 1} (1 - \vartheta)^{n_T + \beta} \; d\vartheta. \end{align} Integrating the former by parts (and observing that the boundary term is \(0\) because it vanishes at both \(0\) and \(1\)) gives \begin{align} \hat{\mathcal{P}}(1) =& \frac{1}{Z} \int_0^1 \left( n_H + \alpha \right)\vartheta^{n_H + \alpha - 1} \cdot \frac{1}{n_T + \beta} (1 - \vartheta)^{n_T + \beta} \; d\vartheta \\=& \frac{n_H + \alpha}{n_T + \beta} \hat{\mathcal{P}}(0) \\=& \frac{n_H + \alpha}{n_T + \beta} \left( 1 - \hat{\mathcal{P}}(1)\right), \end{align} and so \[ \hat{\mathcal{P}}(1) = \frac{n_H + \alpha}{n_H + n_T + \alpha + \beta}. \] Effectively, this says that the Bayesian posterior for a fresh example drawn from the distribution is the same as the distribution of an example selected at random from the training set with the \(\alpha + \beta\) extra "hallucinated" examples added.
As always the differences are subtle. In MLE we maximize \(\log\left(\mathbf{P}(\mathcal{D};\theta)\right)\) in MAP we maximize \(\log\left(\mathbf{P}(\mathcal{D}|\theta)\right)+\log\left(\mathbf{P}(\theta)\right)\). So essentially in MAP we only add the term \(\log\left(\mathbf{P}(\theta)\right)\) to our optimization. This term is independent of the data and penalizes if the parameters, \(\theta\) deviate too much from what we believe is reasonable. We will later revisit this as a form of regularization, where \(\log\left(\mathbf{P}(\theta)\right)\) will be interpreted as a measure of classifier complexity.